Anti-LC3 pAb (Polyclonal Antibody)
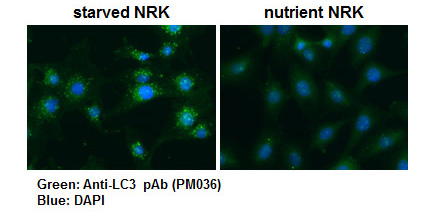
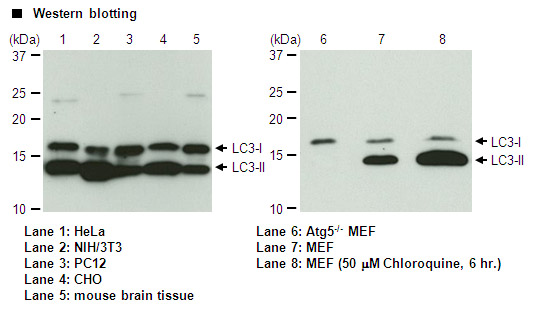
This LC3 antibody is validated for multiple applications (WB, IHC, ICC and IP) and isoform detection. PM036 has more than 1,000 citations where it used to generate data showing autophagy in research papers. (Saitoh T et al. Loss of the autophagy protein Atg16L1 enhances endotoxin-induced IL-1beta production. Nature 456, 264-8 (2008)). This is a polyclonal antibody that is raised in rabbit and is reactive with human, hamster, mouse, rat.
| Target | LC3 |
|---|---|
| Product Type | Antibody |
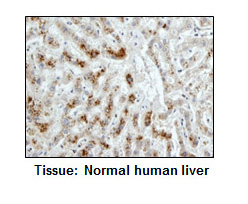
| Application | FCM, ICC, IHC, IP, WB |
| Clonality | Polyclonal |
| Conjugate | Unlabeled |
| Isotype | IgG |
| Immunogen | Recombinant human LC3 (MAP1LC3B :1-120 a.a.) |
| Host Species | Rabbit |
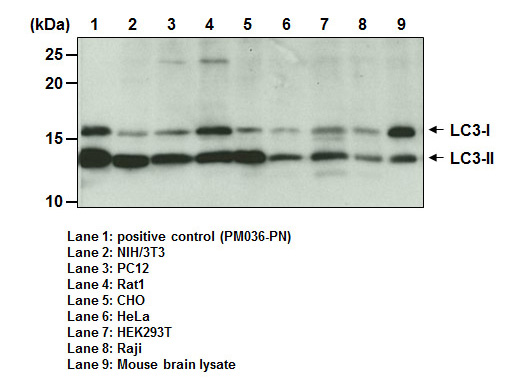
| Species Reactivity | Human, Hamster, Mouse, Rat |
| Formulation | PBS/50% glycerol, pH 7.2 |
| Research Area | Autophagy |
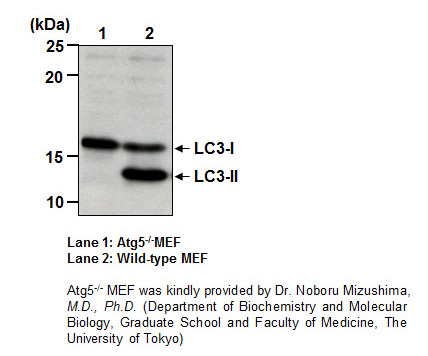
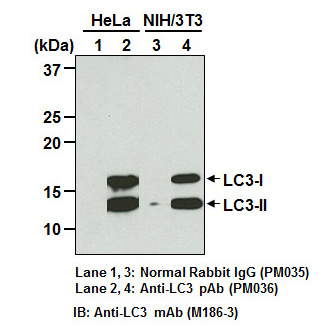
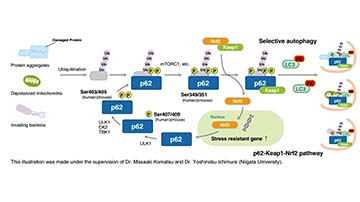
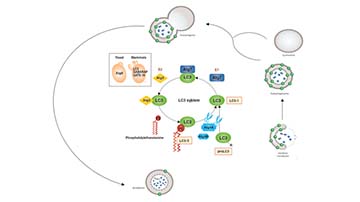
| Description/Background | Macroautophagy mediates the bulk degradation of cytoplasmic components. These components are delivered to lysosomes via autophagosomes. The microtubule-associated protein 1 light chain 3 (LC3), a homologue of yeast Atg8 (Aut7/Apg8), localizes to autophagosomal membranes after post-translational modifications. The C-terminal fragment of LC3 is cleaved immediately following synthesis to yield a cytosolic form called LC3-I. A subpopulation of LC3-I is further converted to an autophagosome-associating form, LC3-II. This antibody can detect forms of LC3 (MAP1LC3A, B, C).</p> |
| Storage Temperature | -20°C |
| Protocols | FCM, ICC, IHC, IP, WB |
| Regulatory Statement | For Research Use Only. Not for use in diagnostic procedures. |